Restriction Enzyme Cleavage of DNA
Familiarization of Gel Electrophoresis and its Uses
Laboratory 5, AP Biology
Abstract. This lab consisted of exploring gel electrophoresis and its use to identify the different DNA pieces that result from a restriction endonuclease digest. The initial preparation for the lab was conducted on a separate day, as to allow for the gel to harden and settle. After gathering sufficient data, we then proceeded to graph the length of the strands from HindIII, EcoRI, and Lambda Uncut.
Introduction
The main discovery aspect of this experiment was the use of restriction enzymes (or endonucleases) and how they specifically cleave or splice DNA in specific locations. Restriction enzymes have the ability to behave in that manner because of short sequences of nucleotides called recognition sequences, this sequence aligns specifically with that of the enzyme's allowing it to be recognized, then cut by the restriction enzyme. The ability of cutting only in specific locations has proven to be beneficial in bacteria, as the restriction enzymes splice invading viruses, but the bacteria itself is safe from any splicing because of a slight modification where the restriction enzyme would normally cut. It is those simple but specific readings that make restriction enzymes function the way they do (LabBench 2014).
Methods
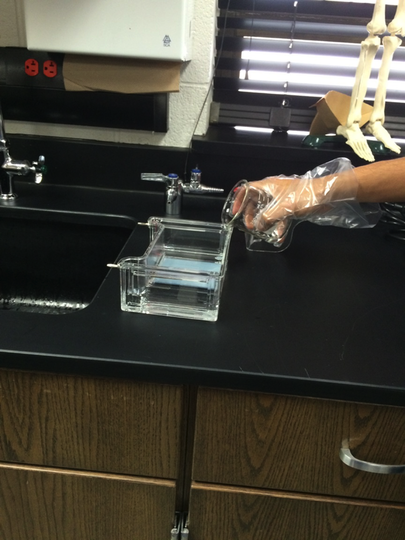
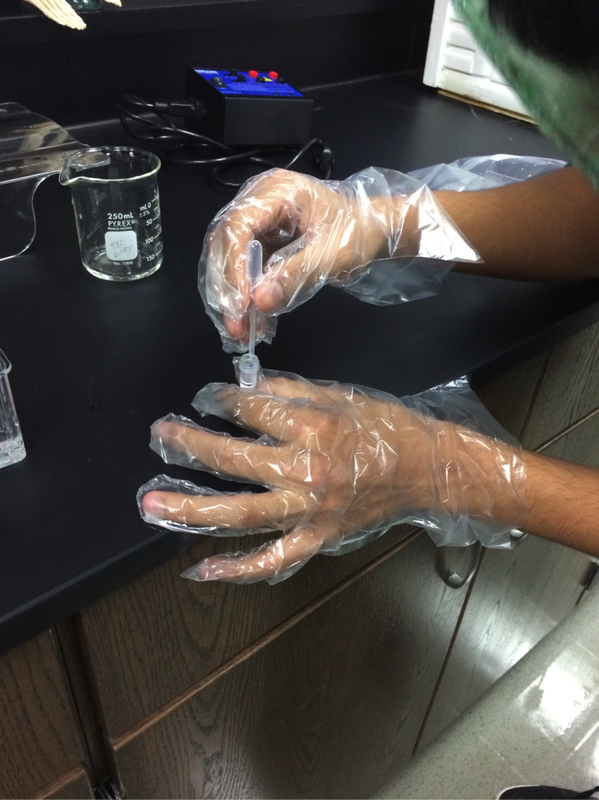
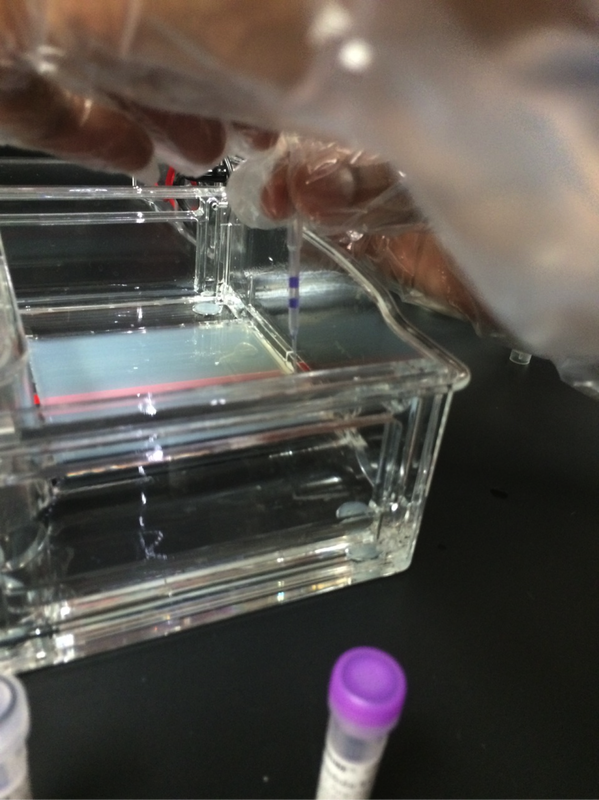
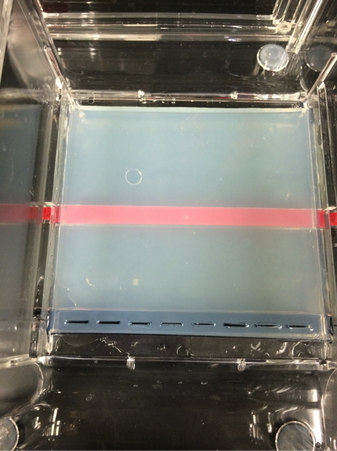
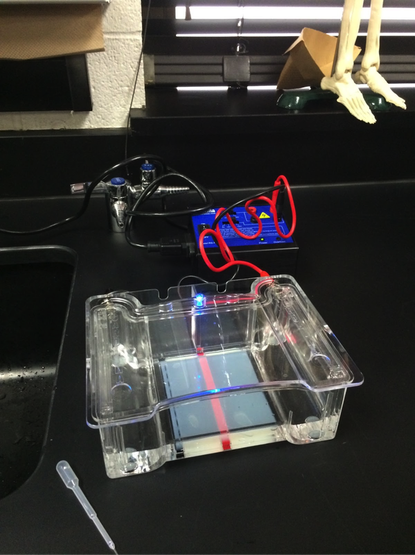
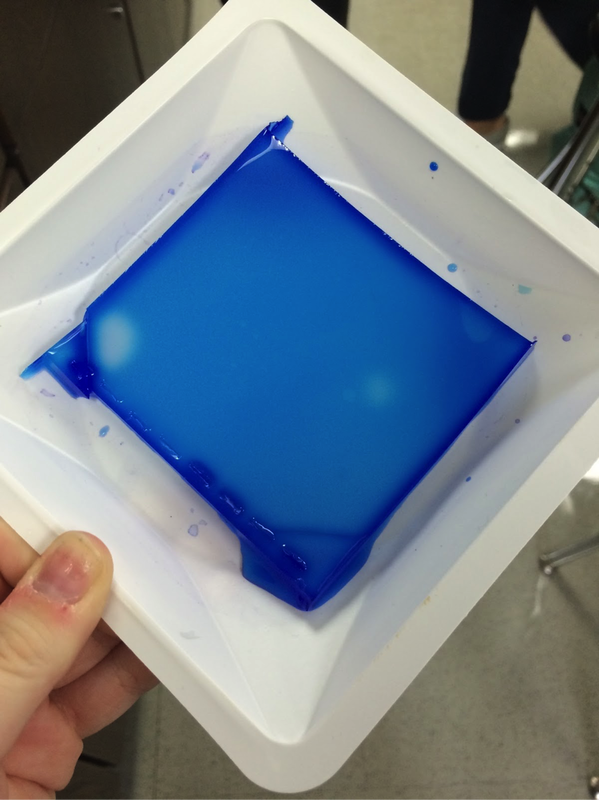
Our experiment was designed by Pearson LabBench. First, we made a gel base using agarose, a seaweed extract. Once solidified, we submerged the gel inside a liquid buffer (Figure 5). We then injected samples of E. Coli DNA (with the restriction enzymes of HindIII, EcoRI, and one without enzymes) into wells left by the solidifying process (Figures 7-11). Next, we applied steady current into the buffer and thus into the gel as well (Figure 12). Once the current had been flowing for approximately 30 minutes, we removed the gel and stored it until the next day, stained with blue dye. The next day, we removed the gel from storage and viewed our results above a light source (LabBench 2014) (Figures 13, 14).
Results
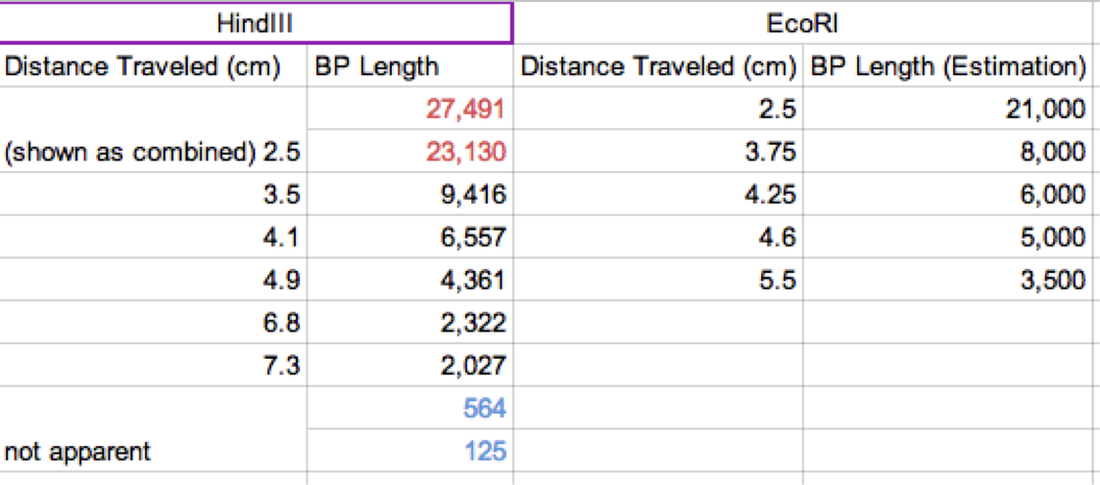
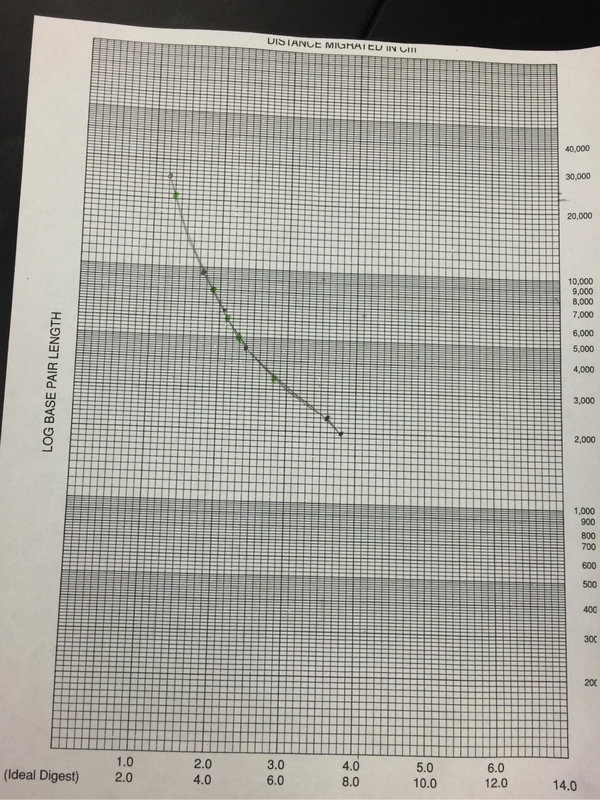
Our results were inconclusive. Perhaps this was due the unsteady hands of ours that shook the DNA upon ejection. Perhaps it was from our novice ability in dealing with miniscule molecules. Whatever the circumstance, we did not yield any results. Due to this, we used the provided results given by Pearson LabBench. In our table, we were given values of the BP length (or pase pair length). We then measured out the given traveling distances to find the distance traveled for each sample (Figure 1). We then graphed using logarithmic graphing paper (Figure 15). From this we were able to extrapolate information for the second column base pair length. Figure 1 shows our results in table format (Carroll, Chowdhury, Fox, Rodriguez, Thomas. (2014)/LabBench 2014).
Discussion
This lab was conducted in order to explore electrophoresis and identify DNA however we were unable to ascertain adequate results. This is most likely not be an error with the experiment but instead a human error done by us conducting the experiment. Due to our inconclusive results, we are unable to articulate whether the findings were in accord with our hypothesis. Additional experiments must be conducted in order to receive accurate results.
However, the research conducted for our lab provided us with valuable knowledge. A restriction enzyme is an enzyme that cuts DNA at a specific point marked by base pair points. Each restriction enzyme is capable of cutting one specific combination of base pairs. We excluded the points plotted for the 27,491/23,130 bp doublet in the standard curve because they require a lower percentage gel than those shorter fragments. This tells us that in order to get accurate results with larger fragments, a 0.8% is much too high. If we wanted to separate DNA fragments that are 25,000 bp, 21,000 bp, 10,000 bp, and 6,000 bp we should use agarose gel around 0.5% to 1%. If we were to separate DNA fragments that are 1,000 bp, 500 bp, or 100 bp, we should use an agarose gel of about 2.5%. If our samples were to have moved in the opposite direction than that which we were expecting, we most likely hooked the negative and positive wires to the wrong sides of the gel tank. The positive and negative wires from the power source are hooked up to each end of the gel tank. The gel as well as the buffer solution is inside the tank, and since DNA is negatively charged, it will be propelled from the negative side towards the positive side. The gel allows small DNA fragments travel quickly while the larger fragments have a much harder time. If we stop the gel at any given time and the band has gone up it, then we likely hooked up the +ve and -ve wires to the wrong sides of the gel tank.
Conclusion
In this experiment, we learned the function and overall significance of electrophoresis, and discovered it is is used to separate macromolecules like DNA or RNA by size or proteins by charge. We used RFLP (Restriction fragment length polymorphism) analysis, a laboratory method in which DNA sample is broken down into fragments by restriction enzymes, and the resulting fragments are separated based on their lengths by gel electrophoresis. This experiment also heightened our understanding of ligase, the “glue” that binds the two smaller enzymes into one single structure. Though the results were inconclusive, conducting this experiment allowed us to explore and understand gel electrophoresis as well as restriction enzymes on a deeper level.
Citations
Carroll, Chowdhury, Fox, Rodriguez, Thomas. (2014). Electrophoresis Student Input.
LabBench. (n.d.). Retrieved December 9, 2014, from http://www.phschool.com/science/biology_place/labbench/lab6/intro.html
Photos and Charts:
Familiarization of Gel Electrophoresis and its Uses
Laboratory 5, AP Biology
Abstract. This lab consisted of exploring gel electrophoresis and its use to identify the different DNA pieces that result from a restriction endonuclease digest. The initial preparation for the lab was conducted on a separate day, as to allow for the gel to harden and settle. After gathering sufficient data, we then proceeded to graph the length of the strands from HindIII, EcoRI, and Lambda Uncut.
Introduction
The main discovery aspect of this experiment was the use of restriction enzymes (or endonucleases) and how they specifically cleave or splice DNA in specific locations. Restriction enzymes have the ability to behave in that manner because of short sequences of nucleotides called recognition sequences, this sequence aligns specifically with that of the enzyme's allowing it to be recognized, then cut by the restriction enzyme. The ability of cutting only in specific locations has proven to be beneficial in bacteria, as the restriction enzymes splice invading viruses, but the bacteria itself is safe from any splicing because of a slight modification where the restriction enzyme would normally cut. It is those simple but specific readings that make restriction enzymes function the way they do (LabBench 2014).
Methods
Our experiment was designed by Pearson LabBench. First, we made a gel base using agarose, a seaweed extract. Once solidified, we submerged the gel inside a liquid buffer (Figure 5). We then injected samples of E. Coli DNA (with the restriction enzymes of HindIII, EcoRI, and one without enzymes) into wells left by the solidifying process (Figures 7-11). Next, we applied steady current into the buffer and thus into the gel as well (Figure 12). Once the current had been flowing for approximately 30 minutes, we removed the gel and stored it until the next day, stained with blue dye. The next day, we removed the gel from storage and viewed our results above a light source (LabBench 2014) (Figures 13, 14).
Results
Our results were inconclusive. Perhaps this was due the unsteady hands of ours that shook the DNA upon ejection. Perhaps it was from our novice ability in dealing with miniscule molecules. Whatever the circumstance, we did not yield any results. Due to this, we used the provided results given by Pearson LabBench. In our table, we were given values of the BP length (or pase pair length). We then measured out the given traveling distances to find the distance traveled for each sample (Figure 1). We then graphed using logarithmic graphing paper (Figure 15). From this we were able to extrapolate information for the second column base pair length. Figure 1 shows our results in table format (Carroll, Chowdhury, Fox, Rodriguez, Thomas. (2014)/LabBench 2014).
Discussion
This lab was conducted in order to explore electrophoresis and identify DNA however we were unable to ascertain adequate results. This is most likely not be an error with the experiment but instead a human error done by us conducting the experiment. Due to our inconclusive results, we are unable to articulate whether the findings were in accord with our hypothesis. Additional experiments must be conducted in order to receive accurate results.
However, the research conducted for our lab provided us with valuable knowledge. A restriction enzyme is an enzyme that cuts DNA at a specific point marked by base pair points. Each restriction enzyme is capable of cutting one specific combination of base pairs. We excluded the points plotted for the 27,491/23,130 bp doublet in the standard curve because they require a lower percentage gel than those shorter fragments. This tells us that in order to get accurate results with larger fragments, a 0.8% is much too high. If we wanted to separate DNA fragments that are 25,000 bp, 21,000 bp, 10,000 bp, and 6,000 bp we should use agarose gel around 0.5% to 1%. If we were to separate DNA fragments that are 1,000 bp, 500 bp, or 100 bp, we should use an agarose gel of about 2.5%. If our samples were to have moved in the opposite direction than that which we were expecting, we most likely hooked the negative and positive wires to the wrong sides of the gel tank. The positive and negative wires from the power source are hooked up to each end of the gel tank. The gel as well as the buffer solution is inside the tank, and since DNA is negatively charged, it will be propelled from the negative side towards the positive side. The gel allows small DNA fragments travel quickly while the larger fragments have a much harder time. If we stop the gel at any given time and the band has gone up it, then we likely hooked up the +ve and -ve wires to the wrong sides of the gel tank.
Conclusion
In this experiment, we learned the function and overall significance of electrophoresis, and discovered it is is used to separate macromolecules like DNA or RNA by size or proteins by charge. We used RFLP (Restriction fragment length polymorphism) analysis, a laboratory method in which DNA sample is broken down into fragments by restriction enzymes, and the resulting fragments are separated based on their lengths by gel electrophoresis. This experiment also heightened our understanding of ligase, the “glue” that binds the two smaller enzymes into one single structure. Though the results were inconclusive, conducting this experiment allowed us to explore and understand gel electrophoresis as well as restriction enzymes on a deeper level.
Citations
Carroll, Chowdhury, Fox, Rodriguez, Thomas. (2014). Electrophoresis Student Input.
LabBench. (n.d.). Retrieved December 9, 2014, from http://www.phschool.com/science/biology_place/labbench/lab6/intro.html
Photos and Charts: